Fluoroquinolones: Mechanisms of Action and Resistance
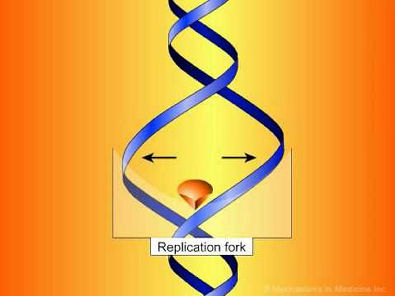

Developed and produced by http://www.MechanismsinMedicine.com Animation Description: In this animation, we demonstrate the biology of DNA replication leading to bacterial cell division in a gram positive bacterium, such as S. pneumoniae. The DNA is shown as a circular double strand within the bacterial cell. Like the DNA of all living organisms, it contains the unique genetic code for all of the proteins required for bacterial survival. Bacteria replicate by a process known as binary fission whereby one bacterium separates into 2 new daughter cells. However, before this can occur, the bacterium must make an identical copy of its complete circular DNA. DNA replication requires that the two strands of DNA separate so that the genetic code of the bacterium can be read and a new complimentary strand can be created for each of the original strands. To accomplish this, various enzymes known as helicases break the hydrogen bonds between the bases in the two DNA strands, unwind the strands from each other, and stabilize the exposed single strands, preventing them from joining back together. The points at which the two strands of DNA separate to allow replication of DNA are known as replication forks. The enzymes DNA polymerase then move along each strand of DNA, behind each replication fork synthesizing new DNA strands (in red) complementary to the original ones. As the replication forks move forward, positive superhelical twists in the DNA begin to accumulate ahead of them. In order for DNA replication to continue, these superhelical twists must be removed. The bacterial enzyme, DNA gyrase, which is also known as topoisomerase II, is responsible for removing the positive superhelical twists so that DNA replications can procede. DNA gyrase is an essential bacterial enzyme composed of two A and two B subunits which are products of the gyrA and gyrB genes. This enzyme has other important functions which affect the initiation of DNA replication and transcription of many genes. With the combined involvement of these enzymes, an entire duplicate copy of the bacterial genome is produced as the 2 replication forks move in opposite directions around the circular DNA genome. Eventually, as the 2 replication forks meet, 2 new complete chromosomes have been made, each consisting of 1 old and 1 new strand of DNA. This is referred to as semi-conservative replication. In order to allow the 2 new interlinked chromosomes to come apart, another bacterial enzyme is needed which is known as topoisomerase IV. This enzyme is structurally related to DNA gyrase and is coded for by the parC and parE genes. Topoisomerase IV allows for the 2 new inter-linked chromosomes to separate so that they can be segregated into 2 new daughter bacterial cells. Complete separation of bacterial cells Fluoroquinolones. First mechanism of action -- inhibition of DNA gyrase. Fluoroquinolones act by inhibiting the activity of both the DNA gyrase and the topoisomerase IV enzymes. For most gram negative bacteria, DNA gyrase is the primary fluoroquinolone target. Fluoroquinolones have been shown to bind specifically to the complex of DNA gyrase and DNA rather than to DNA gyrase alone. As a result of this binding, quinolones appear to stabilize the enzyme-DNA complexes which in turn results in breaks in the DNA that are fatal to the bacterium. A second mechanism of fluoroquinolone action is shown here. With some exceptions, topoisomerase IV is the primary target of fluoroquinolone action in most gram positive bacteria such as Staphylococci and Streptococci, with DNA gyrase being a secondary target. The separation of 2 new interlinked daughter strands of circular DNA is disrupted. The final result on the bacteria, however, is the same. Bacterial replication is disrupted and the bacterium breaks apart. The relative potency of different fluoroquinolone antibiotics (and thus their spectrum of activity) is dependent in part on their affinity for either DNA gyrase or topoisomerase IV or both. One of the most common mechanisms by which bacteria acquire resistance to fluoroquinolones is by spontaneously occurring mutations in chromosomal genes that alter the target enzymes -- DNA gyrase and topoisomerase IV or both. The frequency with which these spontaneous mutations occurs may be in the range of 10-6. The effect of mutations on the activity of an individual fluoroquinolone will vary depending on the number of mutations, the location of the mutations and which target enzyme is affected. If a mutation occurs (either in the gyrA or gyrB gene) that alters DNA gyrase and results in a reduced affinity of the fluoroquinolone antibiotic for this enzyme, the organism will become resistant. View animation to read more.
Compartilhar